"DIY bio" entries

Announcing BioCoder issue 8
BioCoder 8: neuroscience, robotics, gene editing, microbiome sequence analysis, and more.
Download a free copy of the new edition of BioCoder, our newsletter covering the biological revolution.
We are thrilled to announce the eighth issue of BioCoder. This marks two years of diverse, educational, and cutting-edge content, and this issue is no exception. Highlighted in this issue are technologies and tools that span neuroscience, diagnostics, robotics, gene editing, microbiome sequence analysis, and more.Glen Martin interviewed Tim Marzullo, co-founder of Backyard Brains, to learn more about how their easy-to-use kits, like the RoboRoach, demonstrate how nervous systems work.
Marc DeJohn from Biomeme discusses their smartphone diagnostics technology for on-site gene detection of disease, biothreat targets, and much more.
Daniel Modulevsky, Charles Cuerrier, and Andrew Pelling from Pelling Lab at the University of Ottawa discuss different types of open source biomaterials for regenerative medicine and their use of de-cellularized apple tissue to generate 3D scaffolds for cells. If you follow their tutorial, you can do it, too!
aBioBot, highlighted by co-founder Raghu Machiraju, is a device that uses visual sensing and feedback to perform encodable laboratory tasks. Machiraju argues that “progress in biotechnology will come from the use of open user interfaces and open-specification middleware to drive and operate flexible robotic platforms.” Read more…

The revolution in biology is here, now
Biological products have always seemed far off. BioFabricate showed that they're not.

Installation view of The Living’s Hy-Fi, the winning project of The Museum of Modern Art and MoMA PS1’s 2014 Young Architects Program. Photograph by Kris Graves.
The products discussed at BioFabricate aren’t what I thought they’d be. I’ve been asked plenty of times (and I’ve asked plenty of times), “what’s the killer product for synthetic biology?” BioFabricate convinced me that that’s the wrong question. We may never have some kind of biological iPod. That isn’t the right way to think.
What I saw, instead, was real products that you might never notice. Bricks made from sand that are held together by microbes designed to excrete the binder. Bricks and packing material made from fungus (mycelium). Plastic excreted by bacteria that consume waste methane from sewage plants. You wouldn’t know, or care, whether your plastic Lego blocks are made from petroleum or from bacteria, but there’s a huge ecological difference. You wouldn’t know, or care, what goes into the bricks used in the new school, but the construction boom in Dubai has made a desert city one of the world’s largest importers of sand. Wind-blown desert sand isn’t useful for industrial brickmaking, but the microbes have no problem making bricks from it. And you may not care whether packing materials are made of styrofoam or fungus, but I despise the bag of packing peanuts sitting in my basement waiting to be recycled. You can throw the fungal packing material into the garden, and it will decompose into fertilizer in a couple of days. Read more…

A programming language for biology
Antha is a high-level, open source language for specifying biological workflows.
Editor’s note: This is part of our investigation into synthetic biology and bioengineering. For more, download the new BioCoder Fall 2014 issue here.
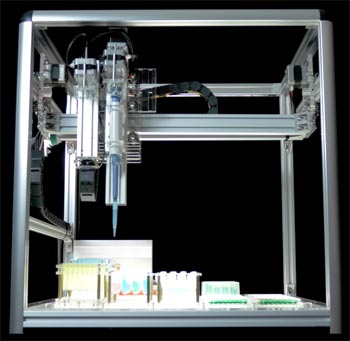

The OT.One liquid handling robot, photo courtesy of OpenTrons.
A programming language for scientific experiments is important for many reasons. Most simply, a scientist in training spends many, many hours of time learning how to do lab work. That sounds impressive, but it really means moving very small amounts of liquid from one place to another. Thousands of times a day, thousands of days in preparation for a career. It’s boring, dull, and necessary work, and something that can be automated. Biologists should spend most of their time thinking about biology, designing experiments, and analyzing results — not handling liquids. Read more…

The future of food
We could soon have lab-grown hamburgers, not in the $300,000 range but in the $10 range — would you eat one?
Editor’s note: this is an excerpt from the latest edition of BioCoder; it is republished here with permission. Get your free copy of BioCoder Fall 2014 here.

Screenshot from the Real Vegan Cheese Indiegogo campaign video.
That was the call I got from a scientist entrepreneur friend of mine, John Schloendorn, the CEO of Gene and Cell Technologies. He’d been working on potential regenerative medicine therapies and tinkering with bioreactors to grow human cell lines. He left the lab for the weekend, and then something went wrong with one of his bioreactors: something got stuck in it.
“So, I was wondering what happened with my bioreactor and how this big chunk of plastic had gotten in there and ruined my cytokine production run. I was pulling it out, and I thought it was was weird because it was floppy. I threw it in the garbage. A little later, after thinking about it, I realized it wasn’t plastic and pulled it out of the garbage.” Read more…

A glowing trend
Glowing plants disrupt the GMO narrative.
 Editor’s note: this is an excerpt from the latest edition of BioCoder; it is republished here with permission. Get your free copy of BioCoder Fall 2014 here.
Editor’s note: this is an excerpt from the latest edition of BioCoder; it is republished here with permission. Get your free copy of BioCoder Fall 2014 here.
Unlike many of his generational peers, Glowing Plant chief scientific officer Kyle Taylor was never put off by genetically modified organism (GMO) crops. On the contrary: Kansas-born and bred, cutting-edge agriculture was as natural to him as the torrid summers and frigid winters of the southern plains.
“GMO corn first hit the market while I was still in high school,” says Taylor, “and I have to admit I was fascinated by it. It was Roundup resistant, meaning that it you could spray it with the most commonly used herbicide in commercial agriculture and it would remain unaffected. I found that really profound, a breakthrough.” Read more…

Avoiding the tragedy of the anticommons
We're at the start of a revolution in biology, and it's time for a biological commons.
Editor’s note: this post originally appeared in BioCoder Fall 2014; it is published here with permission. Download a free copy of the new issue here.
A few months ago, I singled out an article in BioCoder about the appearance of open source biology. In his white paper for the Bio-Commons, Rüdiger Trojok writes about a significantly more ambitious vision for open biology: a bio-commons that holds biological intellectual property in trust for the good of all. He also articulates the tragedy of the anticommons, the nightmarish opposite of a bio-commons in which progress is difficult or impossible because “ambiguous and competing intellectual property claims…deter sharing and weaken investment incentives.” Each individual piece of intellectual property is carefully groomed and preserved, but it’s impossible to combine the elements; it’s like a jigsaw puzzle, in which every piece is locked in a separate safe.
We’ve certainly seen the anticommons in computing. Patent trolls are a significant disincentive to innovation; regardless of how weak the patent claim may be, most start-ups just don’t have the money to defend. Could biotechnology head in this direction, too? In the U.S., the Supreme Court has ruled that human genes cannot be patented. But that ruling doesn’t apply to genes from other organisms, and arguably doesn’t apply to modifications of human genes. (I don’t know the status of genetic patents in other countries.) The patentability of biological “inventions” has the potential to make it more difficult to do cutting-edge research in areas like synthetic biology and pharmaceuticals (Trojok points specifically to antibiotics, where research is particularly stagnant). Read more…

BioCoder strikes again
New issue: bioreactors and food production, modeling a worm's brain on a computer and letting it drive a robot, and more.
 The fifth issue of BioCoder is here! We’ve made it into our second year: this revolution is in full swing.
The fifth issue of BioCoder is here! We’ve made it into our second year: this revolution is in full swing.
Rather than talk about how great this issue is (though it is great), I’d like to ask a couple of questions. Post your answers in the comments; we won’t necessarily reply, but we will will read them and take them into account.
- We are always interested in new content, and we’ll take a look at almost anything you send to BioCoder@oreilly.com. In particular, we’d like to get more content from the many biohacker labs, incubators, etc. We know there’s a lot of amazing experimentation out there. But we don’t know what it is; we only see the proverbial tip of the iceberg. What’s the best way to find out what’s going on?
- While we’ve started BioCoder as a quarterly newsletter, that’s a format that already feels a bit stodgy. Would you be better served if BioCoder went web-native? Rather than publishing eight or 10 articles every three months, we’d publish three or four articles a month online. Would that be more useful? Or do you like things the way they are?
And yes, we do have a great issue, with articles about a low-cost MiniPCR, bioreactors and food production, and what happens when you model a worm’s brain on a computer and let it drive a robot. Plus, an interview with Kyle Taylor of the glowing plant project, the next installment in a series on lab safety, and much more. Read more…

Announcing BioCoder issue 4
Inside this issue: implanting evolution, open source biotech consumables, power supplies for systems biology, and more.

The Summer 2014 edition of BioCoder is now available for free download.
We’ve made it to our fourth issue of BioCoder! I’m excited about this issue — it’s the best collection of articles we’ve published so far.
Some of the highlights are:
- Implanting Evolution:
- We spend a lot of time thinking about how to modify other creatures, from microbes on up. What about ourselves? Surgeons already implant pacemakers and insulin pumps into humans. What about other applications? What are the possibilities if you implant NFC and RFID chips?
- Open Source Biotech Consumables:
- One of the biggest problems for grassroots biotech research is the price of ingredients. Some proteins cost thousands of dollars per milligram, hardly affordable by a community lab or a small startup. We can solve that problem with “open source” DNA. This is an exciting development — and a challenge to what we mean by “open source” (I promise to write about that in another post).