- Bank of America Loading up on Bitcoin Patents — The wide-ranging patents cover everything from a “cryptocurrency transaction payment system” which would let users make transactions using cryptocurrency, to risk detection, storing cryptocurrencies offline, and using the blockchain to measure fraudulent activity.
- Vertigo: A Wall-Climbing Robot (Disney Research) — watch the video. YOW! (via David Pescovitz)
- Synthesizing What I Mean — In this paper, we describe SWIM, a tool which suggests code snippets given API-related natural language queries.
- serialusb — this is how you decode USB protocols.
"hacking" entries


Four short links: 30 December 2015
Bitcoin Patents, Wall-Climbing Robot, English 2 Code, and Decoding USB

The Snapchat Leak
4.6 million phone numbers, is one of them yours?
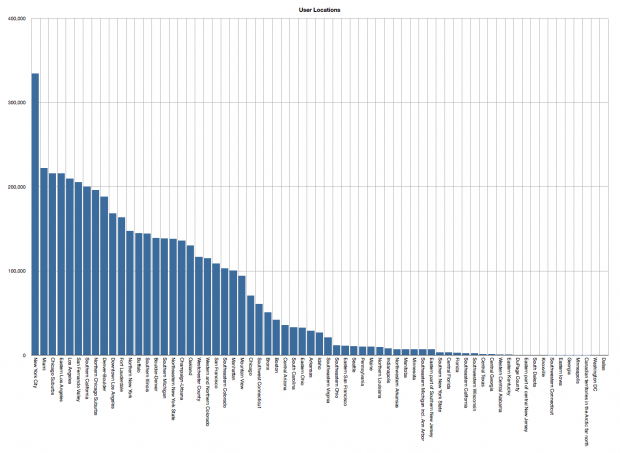
The number of Snapchat users by geographic location. Users are predominately located in New York, San Francisco and the surrounding greater New York and Bay Areas.
While the site crumbled quickly under the weight of so many people trying to get to the leaked data—and has now been suspended—there isn’t really such a thing as putting the genie back in the bottle on the Internet.
Just before Christmas the Australian based Gibson Security published a report highlighting two exploits in the Snapchat API claiming that hackers could easily gain access to users’ personal data. Snapchat dismissed the report, responding that,
Theoretically, if someone were able to upload a huge set of phone numbers, like every number in an area code, or every possible number in the U.S., they could create a database of the results and match usernames to phone numbers that way.
Adding that they had various “safeguards” in place to make it difficult to do that. However it seems likely that—despite being explicitly mentioned in the initial report four months previously—none of these safeguards included rate limiting requests to their server, because someone seems to have taken them up on their offer.


Four short links: 28 November 2013
Data Tool, Arduino-like Board, Learn to Code via Videogames, and Creative Commons 4.0 Out
-
OpenRefine — (edited: 7 Dec 2013)
Google abandonedGoogle bought Freebase’s GridWorks, turned it into the excellent Refine tool for working with data sets, now picked up and developed by open source community. - Intel’s Arduino-Compatible Board — launched at MakerFaire Rome. (via Wired UK)
- Game Maven — learn to code by writing casual videogames. (via Greg Linden)
- CC 4.0 Out — The 4.0 licenses are extremely well-suited for use by governments and publishers of public sector information and other data, especially for those in the European Union. This is due to the expansion in license scope, which now covers sui generis database rights that exist there and in a handful of other countries.


Four short links: 18 October 2013
Publishing Bad Research, Reproducing Research, DIY Police Scanner, and Inventing the Future
- Science Not as Self-Correcting As It Thinks (Economist) — REALLY good discussion of the shortcomings in statistical practice by scientists, peer-review failures, and the complexities of experimental procedure and fuzziness of what reproducibility might actually mean.
- Reproducibility Initiative Receives Grant to Validate Landmark Cancer Studies — The key experimental findings from each cancer study will be replicated by experts from the Science Exchange network according to best practices for replication established by the Center for Open Science through the Center’s Open Science Framework, and the impact of the replications will be tracked on Mendeley’s research analytics platform. All of the ultimate publications and data will be freely available online, providing the first publicly available complete dataset of replicated biomedical research and representing a major advancement in the study of reproducibility of research.
- $20 SDR Police Scanner — using software-defined radio to listen to the police band.
- Reimagine the Chemistry Set — $50k prize in contest to design a “chemistry set” type kit that will engage kids as young as 8 and inspire people who are 88. We’re looking for ideas that encourage kids to explore, create, build and question. We’re looking for ideas that honor kids’ curiosity about how things work. Backed by the Moore Foundation and Society for Science and the Public.


Four short links: 8 May 2013
Paperclip Computing, Packet Capture, Offline Wikipedia, and Sensor Databases
- How to Build a Working Digital Computer Out of Paperclips (Evil Mad Scientist) — from a 1967 popular science book showing how to build everything from parts that you might find at a hardware store: items like paper clips, little light bulbs, thread spools, wire, screws, and switches (that can optionally be made from paper clips).
- Moloch (Github) — an open source, large scale IPv4 packet capturing (PCAP), indexing and database system with a simple web GUI.
- Offline Wikipedia Reader (Amazon) — genius, because what Wikipedia needed to be successful was to be read-only. (via BoingBoing)
- Storing and Publishing Sensor Data — rundown of apps and sites for sensor data. (via Pete Warden)

Glowing Plants
I just invested in BioCurious’ Glowing Plants project on Kickstarter. I don’t watch Kickstarter closely, but this is about as fast as I’ve ever seen a project get funded. It went live on Wednesday; in the afternoon, I was backer #170 (more or less), but could see the number of backers ticking upwards constantly as I watched. It was fully funded for $65,000 Thursday; and now sits at 1340 backers (more by the time you read this), with about $84,000 in funding. And there’s a new “stretch” goal: if they make $400,000, they will work on bigger plants, and attempt to create a glowing rose.
Glowing plants are a curiosity; I don’t take seriously the idea that trees will be an alternative to streetlights any time in the near future. But that’s not the point. What’s exciting is that an important and serious biology project can take place in a biohacking lab, rather than in a university or an industrial facility. It’s exciting that this project could potentially become a business; I’m sure there’s a boutique market for glowing roses and living nightlights, if not for biological street lighting. And it’s exciting that we can make new things out of biological parts.
In a conversation last year, Drew Endy said that he wanted synthetic biology to “stay weird,” and that if in ten years, all we had accomplished was create bacteria that made oil from cellulose, we will have failed. Glowing plants are weird. And beautiful. Take a look at their project, fund it, and be the first on your block to have a self-illuminating garden.

Biohacking: The next great wave of innovation
The hacker culture that launched the computing revolution is now taking root in the bio space.
 I’ve been following synthetic biology for the past year or so, and we’re about to see some big changes. Synthetic bio seems to be now where the computer industry was in the late 1970s: still nascent, but about to explode. The hacker culture that drove the development of the personal computer, and that continues to drive technical progress, is forming anew among biohackers.
I’ve been following synthetic biology for the past year or so, and we’re about to see some big changes. Synthetic bio seems to be now where the computer industry was in the late 1970s: still nascent, but about to explode. The hacker culture that drove the development of the personal computer, and that continues to drive technical progress, is forming anew among biohackers.
Computers certainly existed in the ’60s and ’70s, but they were rare, and operated by “professionals” rather than enthusiasts. But an important change took place in the mid-’70s: computing became the domain of amateurs and hobbyists. I read recently that the personal computer revolution started when Steve Wozniak built his own computer in 1975. That’s not quite true, though. Woz was certainly a key player, but he was also part of a club. More important, Silicon Valley’s Homebrew Computer Club wasn’t the only one. At roughly the same time, a friend of mine was building his own computer in a dorm room. And hundreds of people, scattered throughout the U.S. and the rest of the world, were doing the same thing. The revolution wasn’t the result of one person: it was the result of many, all moving in the same direction.
Biohacking has the same kind of momentum. It is breaking out of the confines of academia and research laboratories. There are two significant biohacking hackerspaces in the U.S., GenSpace in New York and BioCurious in California, and more are getting started. Making glowing bacteria (the biological equivalent of “Hello, World!”) is on the curriculum in high school AP bio classes. iGem is an annual competition to build “biological robots.” A grassroots biohacking community is developing, much as it did in computing. That community is transforming biology from a purely professional activity, requiring lab coats, expensive equipment, and other accoutrements, to something that hobbyists and artists can do.
As part of this transformation, the community is navigating the transition from extremely low-level tools to higher-level constructs that are easier to work with. When I first leaned to program on a PDP-8, you had to start the computer by loading a sequence of 13 binary numbers through switches on the front panel. Early microcomputers weren’t much better, but by the time of the first Apples, things had changed. DNA is similar to machine language (except it’s in base four, rather than binary), and in principle hacking DNA isn’t much different from hacking machine code. But synthetic biologists are currently working on the notion of “standard biological parts,” or genetic sequences that enable a cell to perform certain standardized tasks. Standardized parts will give practitioners the ability to work in a “higher level language.” In short, synthetic biology is going through the same transition in usability that computing saw in the ’70s and ’80s. Read more…

Announcing Make's Hardware Innovation Workshop
The Hardware Innovation Workshop will be held May 15-16.
We're announcing the Hardware Innovation Workshop, a new business conference being held during the week of Maker Faire.

Fighting the next mobile war
Recent moves by Apple and Google could ignite the external accessories space.
While you'll likely interact with your smartphone tomorrow in much the same way you interacted with it today, it's quite possible that your smartphone will interact with the world in a very different way. The next mobile war has already begun.
